Purpose
While characterizing phenotypic differences between interfertile Chlamydomonas species 1, we discovered that C. smithii — but not C. reinhardtii — can grow on marine media. In our previous publication, we documented that cells became "larger and rounded" when grown on Marine Broth 2216 agar supplemented with L1 nutrients, but we'd resuspended those cells in water, which could have caused osmotic swelling.
Here, we examined cells grown on solid Marine Broth 2216 agar and imaged in liquid Marine Broth 2216, revealing far more extreme morphological transformations than we initially observed. We're sharing these findings to document this remarkable plasticity and invite feedback on mechanisms, similar phenomena in other systems, and experimental priorities.
Background and goals
In our broader investigation of genotype–phenotype relationships in Chlamydomonas, we've systematically characterized phenotypic differences between two interfertile species: C. reinhardtii and C. smithii 1. One striking difference we discovered was that C. smithii, unlike C. reinhardtii, can grow on high-salt marine media — specifically Marine Broth 2216 (MB) supplemented with L1 nutrients (MB + L1) and Erdschreiber's medium. When we examined cells grown on MB + L1, they appeared enlarged and rounded compared to their typical ellipsoid morphology on standard freshwater media. However, we resuspended cells in water for those initial observations, which could alter turgor pressure and lead to artificial swelling.
Here, we revisited these morphological changes more carefully by growing C. smithii on MB agar (without L1 supplements) and examining cells resuspended in liquid MB rather than water. Our goal was to determine whether the morphological changes we'd observed reflected artifacts of osmotic stress or genuine cellular transformations induced by the medium.
Understanding morphological plasticity — the ability of organisms to alter their physical form in response to environmental cues — provides insights into cellular adaptability with implications for evolution, cell biology, and biotechnology. Several organisms show dramatic morphological changes in specific nutrient conditions: The bacterium Azotobacter vinelandii adopts a distinctive "fungoid" morphology when grown in peptone-supplemented media 2, the alga Micractinium conductrix increases cellular volume over fivefold as a symbiont within Paramecium bursaria 3, and Tetraselmis convolutae loses flagella, cell wall, and eyespot when living as an endosymbiont 4. Our observation that C. smithii undergoes extreme morphological changes in MB — which contains organic nutrients (peptone, yeast extract, and ferric citrate) absent from standard algal media — provides an opportunity to investigate how model microalgae respond morphologically to nutrient-rich conditions.
The approach
We investigated C. smithii morphology across multiple media types using comparative growth assays and high-resolution microscopy.
Strains and maintenance
We used the Chlamydomonas smithii strain CC-1373 (wild-type, mating type +), obtained from the Chlamydomonas Resource Center. We maintained cells on standard tris-acetate-phosphate (TAP) medium with 1.5% agar under 12:12-hour light-dark cycles at ambient temperature. All solid media contained 1.5% agar. Water plates consisted of 1.5% agar suspended in ultrapure water from a Milli-Q EQ 7008 filtration system with a 0.22 µm filter.
Media formulations
We made TAP and Kuhl’s medium in-house, according to the following recipes. We purchased Marine Broth 2216 (Millipore 76448) and Erdschreiber’s medium (UTEX), and list the ingredients specified by their manufacturers here for convenience.
TAP (tris-acetate-phosphate) medium: 375 µM NH₄Cl, 17.5 µM CaCl₂·2H₂O, 20 µM MgSO₄·7H₂O, 6 µM Na₂HPO₄, 4 µM KH₂PO₄, 200 µM Trizma base, 170 µM glacial acetic acid, 0.1% v/v of Hutner's trace elements solution (134 µM Na₂EDTA·2H₂O, 770 µM ZnSO₄·7H₂O, 184 µM H₃BO₃, 26 µM MnCl₂·4H₂O, 18 µM FeSO₄·7H₂O, 7 µM CoCl₂·6H₂O, 5 µM CuSO₄·5H₂O, and 800 nM (NH₄)₆Mo₇O₂₄·4H₂O), suspended in ultrapure water.
Kuhl's medium: 10 mM KNO₃, 4.5 mM NaH₂PO₄·H₂O, 0.5 mM Na₂HPO₄·2H₂O, 1 mM MgSO₄·7H₂O, 0.1 mM CaCl₃·2H₂O, 24.8 µM FeSO₄·7H₂O, 25 µM Na₂EDTA, 987 nM H₃BO₃, 1 µM MnSO₄·H₂O, 1 µM ZnSO₄·7H₂O, 10 nM CuSO₄·5H₂O, 10 nM (NH₄)₆Mo₇O₂₄·4H₂O, suspended in ultrapure water.
Marine Broth 2216 (MB): 5 g/L peptone, 1 g/L yeast extract, 0.41 mM C₆H₅FeO₇ (ferric citrate), 333.1 mM NaCl, 62 mM MgCl₂, 26.9 mM MgSO₄, 16.2 mM CaCl₂, 7.4 mM KCl, 1.9 mM NaHCO₃, 0.67 mM KBr, 0.21 mM SrCl₂, 0.36 mM H₃BO₃, 0.033 mM Na₂SiO₃, 0.06 mM NaF, 0.02 mM NH₄NO₃, and 0.056 mM Na₂HPO₄ in ultrapure water.
Erdschreiber's medium: 2.3 mM NaNO₃, 67 µM Na₂HPO₄·7H₂O, 23.7 µM Na₂EDTA·2H₂O, 4.3 µM FeCl₃·6H₂O, 2.5 µM MnCl₂·4H₂O, 400 nM ZnCl₂, 100 nM CoCl₂·6H₂O, 200 nM Na₂MoO₄·2H₂O, 100 nM cyanocobalamin, and 50 µM HEPES, suspended in synthetic seawater (409 mM NaCl, 53 mM MgCl₂·6H₂O, 28 mM Na₂SO₄, 10 mM CaCl₂·2H₂O, 1 mM KCl, 1 mM NaHCO₃, 1 mM KBr, 0.4 mM SrCl₂·6H₂O, 1 mM H₃BO₃, 3 mM NaOH, and 2 mM NaF [RICCA Chemical Company: R8363000]), supplemented with 50 mL/L "Soil Water: GR+ Medium" (0.05 mM CaCO₃, 2.5% greenhouse soil, suspended in ultrapure water).
A key distinction between MB and Erdschreiber's medium is that MB contains organic nutrients (peptone at 5 g/L, yeast extract at 1 g/L, and ferric citrate), while Erdschreiber's medium contains only inorganic nutrients despite similar salt concentrations (333 mM NaCl in MB versus 409 mM NaCl in Erdschreiber's synthetic seawater base).
Experimental design
We seeded C. smithii from TAP medium onto plates containing TAP, Kuhl's medium, MB, Erdschreiber's medium, or water, each with 1.5% agar. We grew cells at ambient temperature under 12:12-hour light-dark cycles for 43 days. We collected cells from colonies on different media types and immediately transferred them to wells of a #1.5 glass-bottom 96-well black-walled plate (Cellvis, Cat # P96-1.5H-N) containing the corresponding liquid medium on the 40th and 43rd days of growth. This approach allowed us to observe morphology under conditions that matched the growth environment, thereby avoiding osmotic artifacts.
Microscopy
We used a Yokogawa CSU-W1 SoRa spinning disk confocal unit mounted on a Nikon Ti2-E inverted microscope, equipped with an ORCA-Fusion BT sCMOS camera (Hamamatsu) and a LIDA Light Engine (Lumencor) for illumination, controlled with NIS-Elements AR software (v5.42.03). All images were acquired with a Plan Apo λ 40× air objective (NA 0.95). For all confocal fluorescence acquisitions, the 1.5× relay zoom was engaged, yielding an effective pixel size of 0.108 µm. Brightfield images acquired on day 40 without the relay zoom have a pixel size of 0.163 µm.
For subcellular visualization, we stained cells with MitoTracker Orange CMTMRos (ThermoFisher, Cat. # M7510), PKmito ORANGE (Spirochrome; Cytoskeleton, Cat. # CY-SC053), or FM4-64 FX (ThermoFisher, Cat. # F34653) to label mitochondria or plasma membrane, respectively. We used stains at 1× or 10× the manufacturer's recommended concentration, added directly to the medium without a subsequent wash step. All three dyes were imaged using 561 nm excitation with emission collected in the ~595 nm band (TRITC channel). Chloroplast autofluorescence was simultaneously captured using 640 nm excitation with emission collected in the ~700 nm band (Cy5 channel). Both channels were acquired sequentially at each z-plane using the spinning-disk confocal unit. Staining was performed only on Marine Broth cultures as a proof-of-principle experiment; we intended to extend this to additional media conditions but were unable to do so due to contamination of subsequent cultures and a shift in experimental focus.
We acquired fluorescence z-stacks at 35 planes with a 0.6 µm step size (20.4 µm total range), driven by a Piezo Z stage. We acquired one high-resolution z-stack at 0.1 µm steps over 201 planes for a single Marine Broth field. For time-lapse imaging, we used two protocols depending on the biological question: High-speed acquisitions captured cell motility over 30–60 seconds at ~50 ms frame intervals, while a long-term acquisition tracked colony growth over 2 h at 5-minute intervals.
We acquired 2–6 fields of view per media condition at each time point, for a total of 38 static fields across both imaging sessions.
All raw image files are available on BioImage Archive (DOI: 10.6019/S-BIAD2873).
Image processing
Image enhancement for fluorescence time-lapse
We applied a processing workflow to enhance the visibility of subtle cellular structures while preserving temporal dynamics. The workflow included normalization to 8-bit depth, Gaussian smoothing (σ = 2.0) for noise reduction, unsharp masking (enhancement factor = 1.5) to accentuate fine structures, Contrast Limited Adaptive Histogram Equalization (CLAHE, clip limit = 1.5, 8 × 8 tile grid), temporal smoothing using a 3-frame sliding window average, and final spatial Gaussian blur (σ = 0.3). This processing was necessary to visualize the thin appendages extending from cell bodies, which were barely visible in raw images due to their low contrast against the background. We generated comparative videos that display both original and processed images side-by-side to ensure that processing didn't introduce artifacts.
The script we used is available on GitHub as reveal_wisps.py.
Channel registration
We corrected two-channel fluorescence images for chromatic aberration by registering the MitoTracker channel to the transmitted light reference channel. We obtained an initial shift estimate using phase cross-correlation, then manually refined using an interactive overlay. We applied the optimal translation (x, y pixels) to the MitoTracker channel using an affine transformation with bilinear interpolation, preserving original 16-bit intensity values.
The code we used is available on GitHub as realign_channels_clean.ipynb.
Uneven illumination correction
We corrected for uneven background illumination of images acquired with LED illumination (Lumencor Lida) using a flat-field correction approach. For each color channel independently, we generated a background estimate by applying a Gaussian blur (σ = 100 pixels) to the original image. We then divided the original image by this background estimate and rescaled to preserve the original bit depth. This corrects for spatial variations in illumination intensity while preserving the cellular signal.
The script we used is available on GitHub as smoothen_lida_rgb_tifs.py.
We used Python 3.12.0 with NumPy (v1.21.0), OpenCV (v4.5.0), and SciPy (v1.7.0). Visualization used Matplotlib (v3.4.0) and Pillow (v8.3.0), while TIFF operations used tifffile (v2021.11.0) for imaging processing.
All code, including scripts reveal_wisps.py, smoothen_lida_rgb_tifs.py, and the notebook realign_channels_clean.ipynb, is available in our GitHub repo (DOI: 10.5281/zenodo.18637097).
Pub preparation
We assembled figures in Adobe Illustrator. We used Claude (Sonnet 4, Sonnet 4.5) to help write, clean up, comment, and review analysis code, as well as to write text that we then edited, suggest wording ideas, rearrange starting text to fit a template, copy-edit drafts to match Arcadia's style, and clarify and streamline passages. We also used Grammarly Business to suggest wording ideas and selected which small phrases or sentence structure ideas to incorporate. Additionally, we used Gemini (2.5 Pro) to help clarify and streamline text. All AI-generated text and code were reviewed, edited, and selectively incorporated by the authors before use.
The observations
C. smithii adopts dramatically variable morphologies in Marine Broth 2216
When grown on standard freshwater media (TAP, water, or Kuhl's medium) for 40–43 days, C. smithii cells maintained their typical ellipsoid morphology with two anterior flagella; cells were indistinguishable across these media. However, cells grown on MB for the same duration showed extreme morphological transformations when examined in liquid MB (Figure 1, Video 1).
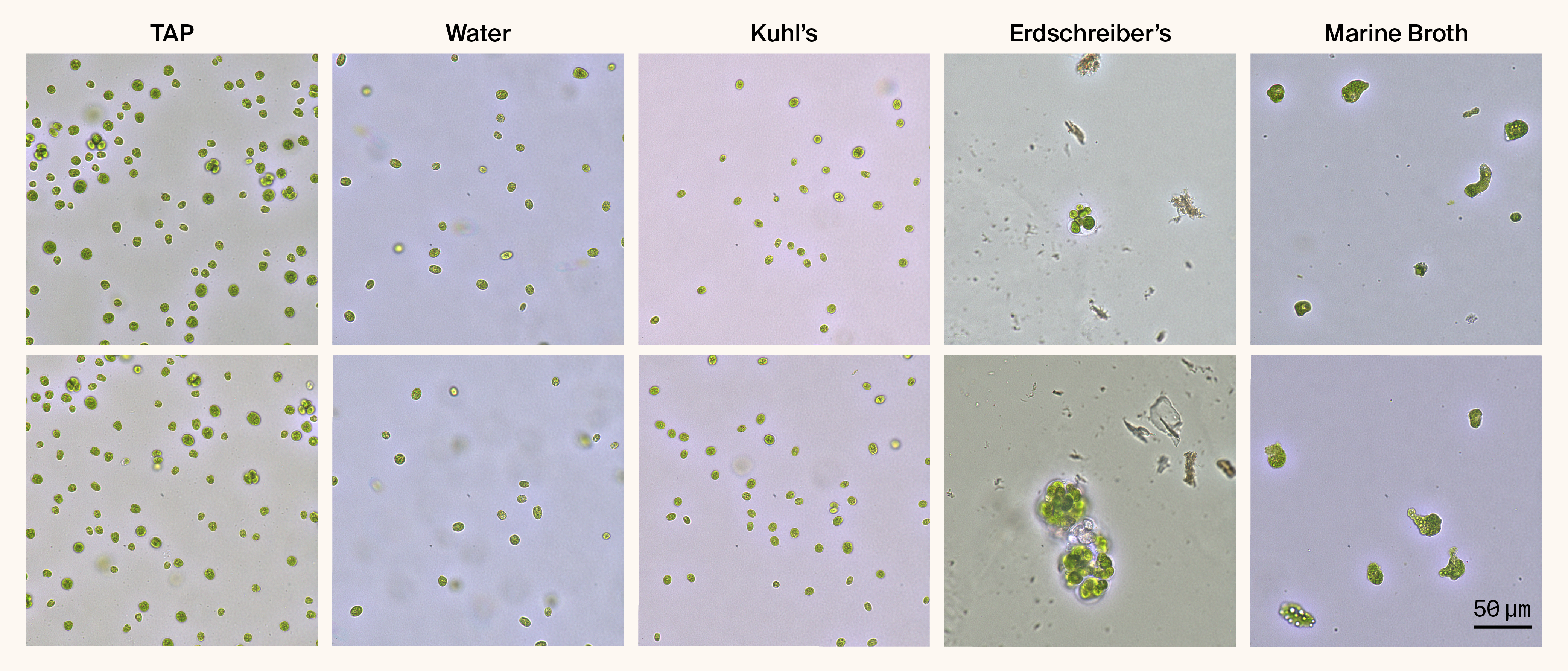
Figure 1. C. smithii exhibits progressive morphological changes across different growth media.
Representative brightfield images of Chlamydomonas smithii grown for 43 days on 1.5% agar plates supplemented with different media, then resuspended in the corresponding liquid medium. TAP, Water, and Kuhl's media maintain normal ellipsoid cell morphology. Erdschreiber's media (409 mM NaCl, no organic nutrients) induces an intermediate phenotype with enlarged, rounded cells and palmelloid-like clusters. MB (333 mM NaCl + 5 g/L peptone + 1 g/L yeast extract) induces extreme morphological transformation, resulting in highly irregular, amorphous cells. Two representative fields are shown for each condition.
These MB-grown cells displayed remarkable variability in size and shape. Unlike the uniform ellipsoid cells we observed in freshwater media, the cells in the present study were dramatically enlarged (appearing 3–5× larger by visual estimation), had highly irregular contours, and exhibited amorphous morphologies with numerous protrusions and indentations. The cells appeared packed with large vacuole-like structures and showed variable coloration, ranging from dark green to pale yellow-green, yet maintained photosynthetic pigmentation, indicating they remained viable. Notably, these cells showed no visible flagella and appeared non-motile. Consistent with our earlier observations, when we exposed them to focused light for an extended period of time, some amorphous cells began to round up 1 (Video 1).
Video 1. C. smithii morphology after 40 days of growth on Marine Broth 2216.
Time-lapse movie of Chlamydomonas smithii grown for 40 days on 1.5% agar plates supplemented with MB, then resuspended in liquid MB. We captured images every 5 min for 100 min (21 frames).
Cells we grew on Erdschreiber's medium, which has similar salt concentrations to MB (409 mM NaCl versus 333 mM NaCl, respectively) but lacks the organic nutrients, appeared larger and more rounded than freshwater-grown cells and frequently formed palmelloid clusters (multiple cells within a common matrix), consistent with a stress response. However, they didn't develop the extreme amorphous morphology or extensive protrusions we observed in MB.
Interestingly, this Erdschreiber's phenotype contrasts with our previous observations 1 where Erdschreiber-grown cells appeared morphologically similar to freshwater-grown cells. The difference might be due to a longer growth duration in the current experiments or batch-to-batch variability in media preparation. This inconsistency itself suggests that subtle environmental factors we haven't controlled for can influence the morphological response.
The fact that similar salt concentrations produce different morphological outcomes in Erdschreiber's versus MB strongly suggests the transformation isn't simply a response to osmotic stress but rather to specific organic nutrients present in MB.
Thin appendages extend from cell bodies
Video 2. Side-by-side comparison of raw and processed imaging reveals wisp-like appendages.
Left: Raw blue channel from RGB brightfield imaging (LIDA illumination).
Right: Same frames after contrast enhancement processing (reveal_wisp.py pipeline: Gaussian smoothing, unsharp masking, CLAHE, temporal smoothing). Multiple thin appendages extending from the MB-grown C. smithii cell body are faintly visible in raw time-lapse but become clearly distinguishable after processing. Processing enhances visibility without introducing artifacts — structures present in raw data are accentuated rather than created. 1-minute time-lapse (1,023 frames at ~58.7 ms intervals) played back at 0.25× speed, blue channel selected for optimal contrast.
We observed one of the most striking features in some MB-grown cells: thin, wisp-like appendages extending from the cell body. These structures were barely visible without image processing but became apparent after contrast enhancement (Video 2). The appendages were extremely thin and highly variable in their length.
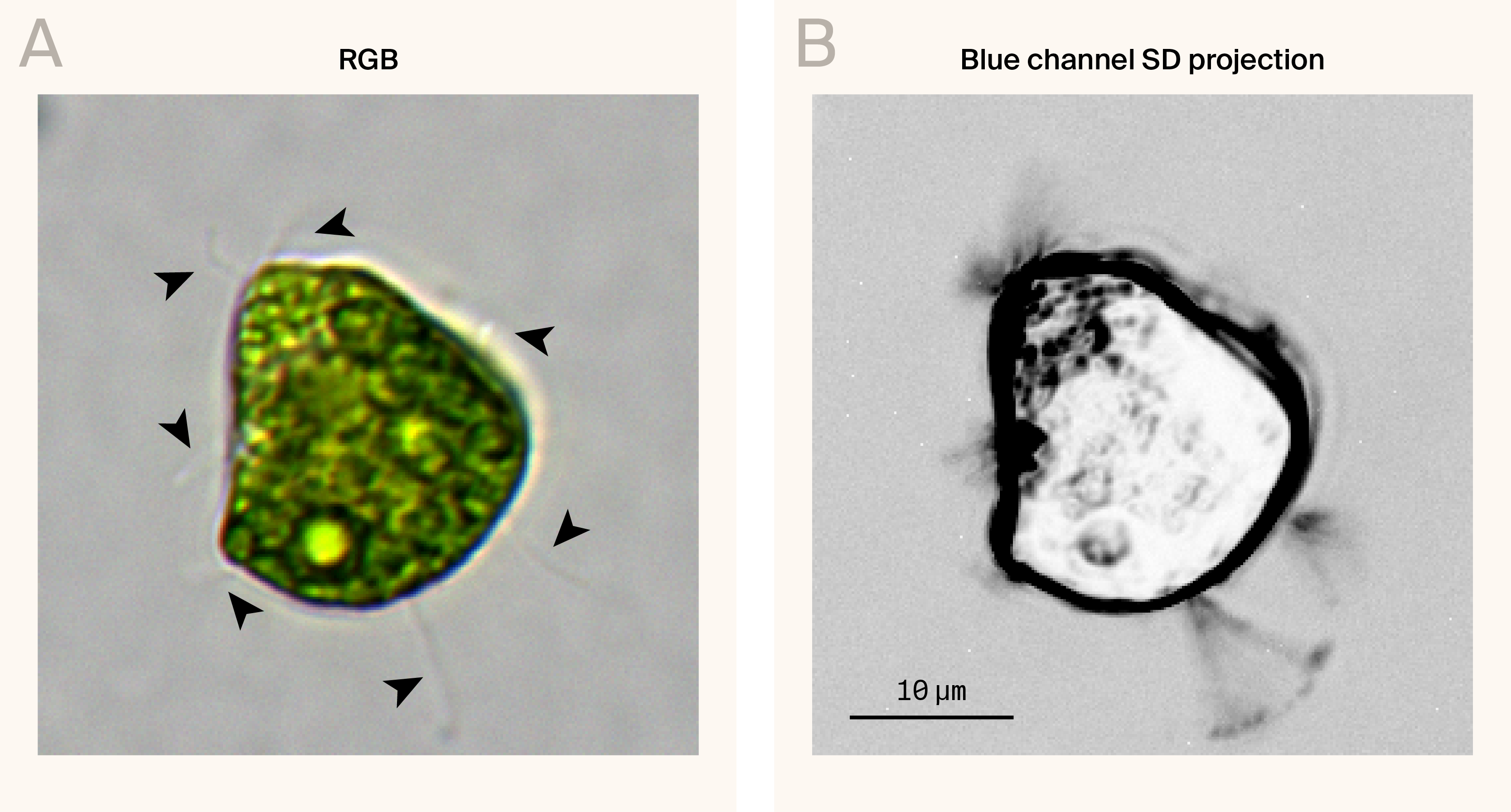
Figure 2. High-magnification image showing wisp-like appendages extending from MB-grown C. smithii.
(A) Reconstructed RGB image of a Chlamydomonas smithii cell grown in MB. Arrowheads indicate wisp-like appendages.
(B) Standard deviation (SD) projection of 280 frames captured at 0.21-second intervals (~4.7 fps) from the blue channel, revealing the dynamic movement of wisp-like structures over a 1-minute period.
Individual cells frequently had multiple appendages extending in different directions, with counts ranging from 0 to 5+ visible appendages per cell in our images. The appendages showed minimal dynamics in our time-lapse videos — they didn't exhibit the active beating characteristic of normal flagella but showed slight passive movement, suggesting they're flexible rather than rigid structures. We display this motility by taking a 1-minute standard deviation projection of a 280-frame time-lapse (Figure 2).
These structures appear to be genuine cellular extensions rather than contaminants based on several observations. Most importantly, when we stained cells with mitochondrial dyes (PKmito ORANGE or MitoTracker Orange), we detected mitochondrial signal within some of these appendages, in at least one case extending nearly 200 µm from the cell body (Figure 3). Additionally, the appendages appear to emerge directly from the cell body rather than being merely adjacent structures, and their occurrence correlates specifically with MB growth conditions rather than occurring randomly, as seen in a manually z-stack through a time-lapse image (Figure 4).
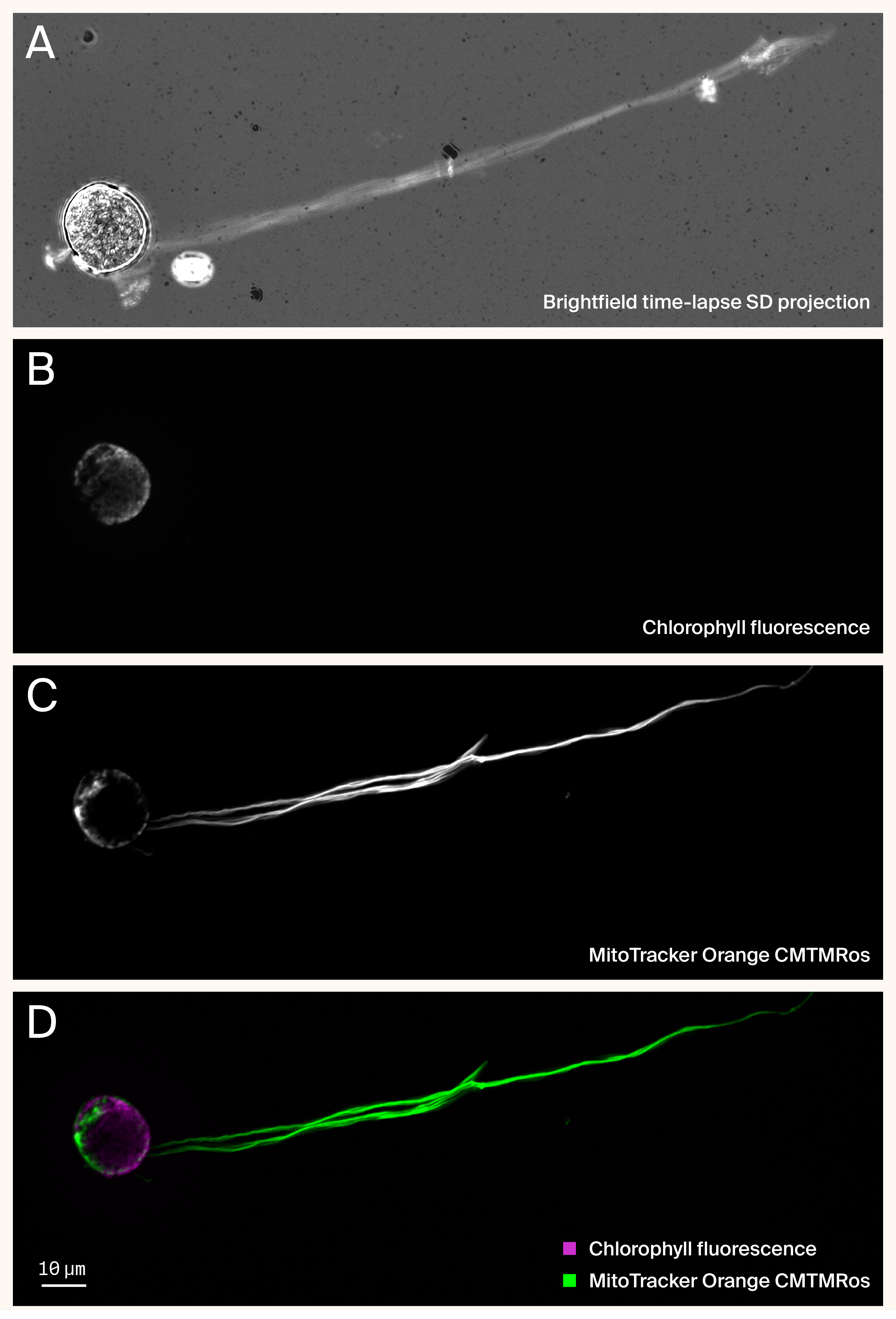
Figure 3. Mitochondrial staining reveals that wisp-like appendages are cellular extensions containing organelles.
(A) Standard deviation projection of a brightfield time-lapse movie (598 frames, 50 ms interval, ~35 s total) showing extensive wisp-like appendages extending from MB-grown C. smithii cell body.
(B) Chlorophyll autofluorescence (magenta) localizes exclusively to the cell body.
(C) Mitochondrial staining (MitoTracker Orange CMTMRos, green) reveals mitochondria within both the cell body and extending throughout the length of the wisp appendage.
(D) Merged image demonstrating spatial separation of chloroplasts (confined to cell body) and mitochondria (present in both cell body and wisp). The presence of mitochondria within wisps suggests these structures are cellular extensions rather than extracellular contaminants.
Whether these are aberrant flagella, novel protrusions, or some other type of cellular extension remains unclear.
Figure 4. Manual focus adjustment reveals three-dimensional complexity of MB-transformed C. smithii cell.
Grayscale brightfield time-lapse with manual z-plane adjustment during acquisition to visualize cellular structure, including vacuole-like compartments and wisp-like appendages extending in different focal planes. The changing focus demonstrates these are genuine three-dimensional cellular structures rather than flat optical artifacts. Acquired at 0.05-second intervals, played at 5 fps (0.25× real-time speed). MB growth conditions (43 days).
Spatial reorganization of chloroplasts and mitochondria
Given the extreme morphological transformations in MB-grown cells, we examined whether internal organellar organization was also altered. When we stained cells with the plasma membrane dye FM 4-64FX, we unsurprisingly observed ring-like structures concentrated around vacuoles, indicating that they're membrane-bound (Figure 5, A).
We also examined the mitochondrial network. Previously, we observed that the mitochondrial network of C. smithii is typically closely associated with the chloroplast, facilitating metabolic cooperation between these organelles 1, 5. However, in several MB-grown cells, we observed striking spatial separation between these organelles. Some morphologically distinct cells showed only sparse punctate mitochondrial staining while chlorophyll autofluorescence remained bright and dominant throughout the cell body (Figure 5, B). In other cells, however, the chloroplast autofluorescence appeared dispersed around the cell periphery with a distinct central zone of exclusion. When we stained these cells with mitochondrial dyes, we detected concentrated mitochondrial signal specifically within this chloroplast-free central zone (Figure 5, C–D).
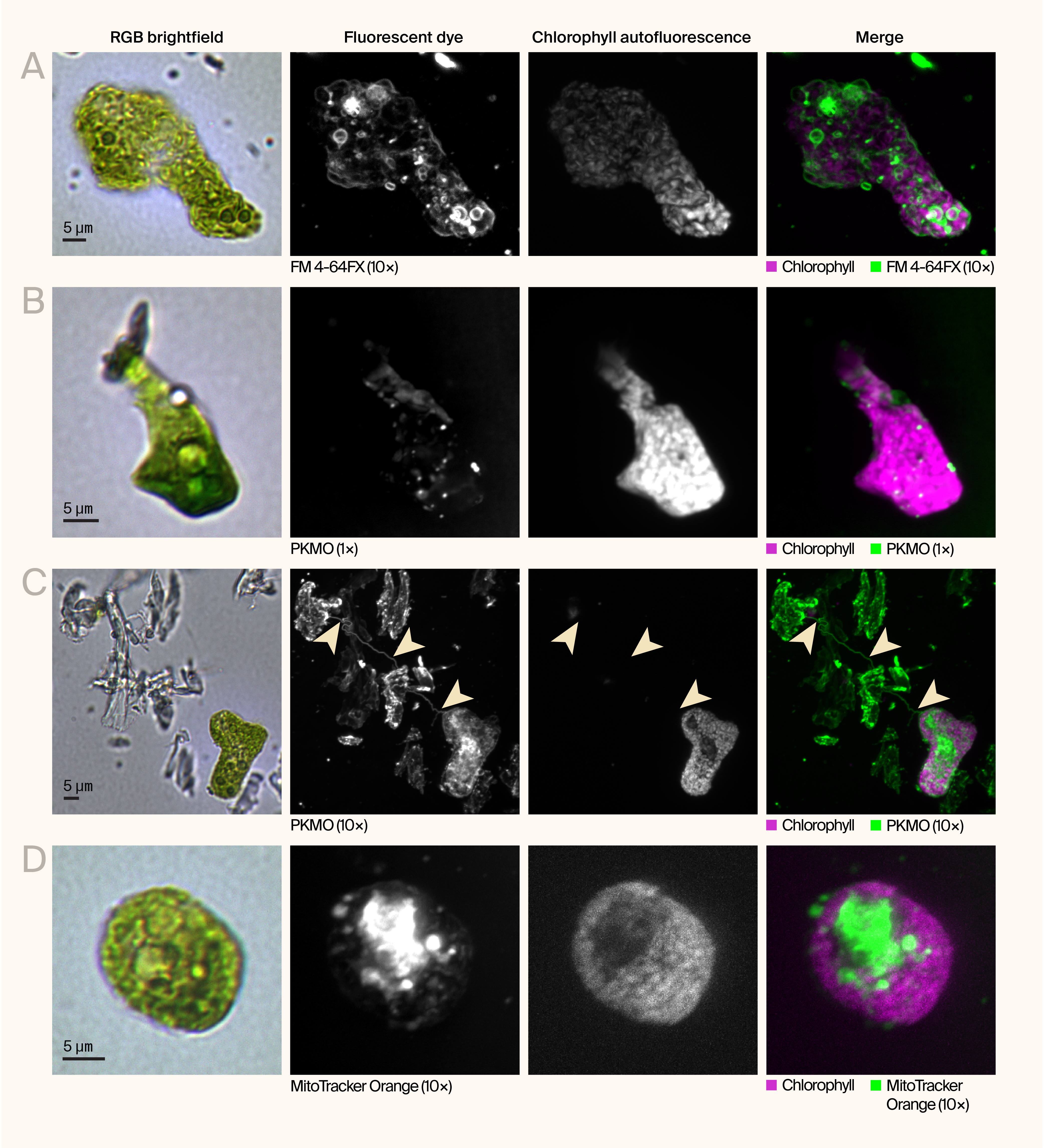
Figure 5. Fluorescent imaging reveals the spatial organization of organelles and membranes in morphologically transformed C. smithii cells.
Representative RGB brightfield images and maximum intensity projections of fluorescently labeled MB-grown C. smithii cells showing diverse morphologies and subcellular organization.
(A) Cell with elongated morphology stained with FM 4-64FX (green, 10× concentration), a lipophilic membrane dye, showing membrane continuity from the cell body through the extension. Chlorophyll autofluorescence (magenta) localizes to the cell body.
(B) Enlarged, irregularly shaped cell stained with PKmito ORANGE (PKMO) (green, 1× concentration) showing mitochondrial distribution. Chlorophyll autofluorescence (magenta) is concentrated in the enlarged cell body.
(C) Field of view showing cellular debris and an intact cell (bottom right) stained with PKmito ORANGE (green, 10× concentration). Arrowheads indicate wisp-like structures persisting in the debris field.
(D) Rounded cell showing more uniform morphology, stained with MitoTracker Orange CMTMRos (green, 10× concentration), with mitochondria and chloroplasts (magenta) distributed throughout the cell body.
This spatial segregation suggests a fundamental reorganization of cellular architecture during the morphological transformation in MB. Whether this reflects altered metabolic requirements, disrupted organellar interactions, or simply physical constraints imposed by the extreme cell shapes remains to be determined. This pattern wasn't universal — some cells showed more typical organellar organization — but it appeared frequently enough to suggest it's a consistent feature of the transformation rather than an occasional aberration.
Access our microscopy data on BioImage Archive.
Thoughts and questions
These observations raise several questions about the mechanisms and significance of this remarkable morphological plasticity:
What components of Marine Broth 2216 trigger the transformation?
MB differs from standard algal media in three major organic components: peptone (5 g/L), yeast extract (1 g/L), and ferric citrate. Based on the observation that Erdschreiber's medium (which lacks these organic nutrients) produces less extreme morphology despite similar salt concentrations, we hypothesize that one or more of these organic components trigger the transformation.
Peptone seems like a likely candidate based on observations that the bacterium Azotobacter vinelandii develops similar extreme shape changes (described as "fungoid" morphology) when grown in peptone-rich media 2. However, we can't rule out contributions from yeast extract or synergistic effects between multiple components. Ferric citrate seems less likely to be solely responsible since Erdschreiber's medium contains iron (as FeCl₃) without producing the same extreme phenotype.
Systematically testing these components individually and in combination would identify what's necessary and sufficient for the transformation.
Reproducibility considerations
We collected the data presented here during a one-week imaging session following 40–43 days of growth on solid media. We've observed similar morphological transformations in MB-grown C. smithii across multiple independent experiments over a two-year period using different batches of cells and media preparations. However, the combination of slow growth and rich medium composition makes this system challenging to work with systematically.
C. smithii grows considerably slower on MB than on standard freshwater media — colonies take 40+ days to become visible on solid MB when seeded from early log phase cultures, compared to 5–10 days on TAP. This extended growth period, combined with MB's high nutrient content (5 g/L peptone, 1 g/L yeast extract), creates ideal conditions for bacterial contamination. We occasionally observed that cultures appearing completely axenic when grown in TAP medium would develop visible bacterial contamination when transferred to MB. This suggests that bacteria may be present but undetectable in standard conditions, then proliferate in the nutrient-rich medium — a well-documented challenge in algal culture work 6. Whether bacterial presence influences the morphological changes remains an open question.
Even in apparently axenic cultures, we observed variable reproducibility. Some batches of MB-grown cells showed more extreme morphologies than others, even with apparently identical growth conditions and duration. We couldn't identify specific variables that controlled this variability, though possibilities include subtle differences in media preparation, fluctuations in light intensity, minor differences in inoculum density, or the presence of undetected contaminants.
What's the adaptive significance?
Why would C. smithii undergo such dramatic morphological changes in MB? Several possibilities:
Heterotrophic adaptation: The changes might facilitate uptake of organic nutrients, shifting from photoautotrophic to mixotrophic metabolism. The loss of flagella and the increase in surface area could reflect a reduced need for motility and an increased capacity for nutrient absorption.
Stress response: Despite abundant nutrients, the combination of high salt and unusual organic nutrients might represent a stressful environment, triggering morphological changes. The palmelloid phenotype in Erdschreiber's supports this interpretation.
Metabolic remodeling: The segregation of chloroplasts to the periphery while mitochondria concentrate centrally might indicate altered metabolic priorities when organic carbon is abundant.
What are the wisp-like appendages?
The thin extensions are particularly puzzling. Possibilities include:
Modified flagella: Perhaps MB components disrupt normal flagellar development, resulting in elongated, dysfunctional flagella
Stress-response structures: Unusual extensions produced under stress conditions
The presence of mitochondria dye uptake within these appendages suggests they're cellular, but their specific identity and function remain unclear.
Next steps
Systematic dissection of Marine Broth 2216 components
The most obvious next experiment is to test individual MB components to identify what triggers the transformation. We plan to supplement TAP medium (which C. smithii grows on well) with:
- Peptone alone (testing multiple concentrations)
- Yeast extract alone
- Ferric citrate alone
- Pairwise combinations of these components
- All three components together (MB composition without salt)
This approach should pinpoint which component(s) are necessary and sufficient while using a medium that C. smithii grows on reliably.
Testing other Chlamydomonas species
Since common C. reinhardtii strains aren't halotolerant and can't grow on full MB 1, we'll test whether it shows similar morphological responses when grown in TAP supplemented with individual MB components or on the freshwater equivalent, Nutrient broth (Millipore N7519). This would help determine whether the capacity for this morphological response is species-specific or if C. reinhardtii simply can't access MB due to salt intolerance.
We want your input
We're sharing these observations before completing systematic component testing because they're striking and unexpected. We'd particularly welcome feedback on:
- Similar observations: Have you seen similar morphological responses in other algae or microorganisms exposed to nutrient-rich media?
- Technical protocols: Do you have protocols for establishing axenic algal cultures, or alternative approaches to test whether bacteria contribute to the phenotype?
- Mechanistic insights: Do you have insights into nutrient-induced morphological changes from other systems?
- Experimental priorities: Given limited resources, which experiments would you prioritize? Component testing? Time course analysis? Ultrastructural characterization?
- Alternative hypotheses: Are there explanations for the wisp-like structures or organellar reorganization we haven't considered?
Acknowledgements
Acknowledgements
We'd like to thank Harper Wood and Hayley Greenough for providing the media that was essential to these observations.